Lac operon
Francois Jacob and Jacques Monad proposed a model for an operon, which consisted of a regulator gene, an operator site containing a regulatory DNA sequence, and one or more structural genes.
The model displayed how stimuli from the environment can promote/inhibit genetic mechanisms which control metabolic events e.g. the presence/absence of glucose and lactose in the lac operon[1]. The lac operon consists of an additional promoter, in front of the regulator gene, the role of which is to ensure the RNA Polymerase binds to the correct transcription initiator.
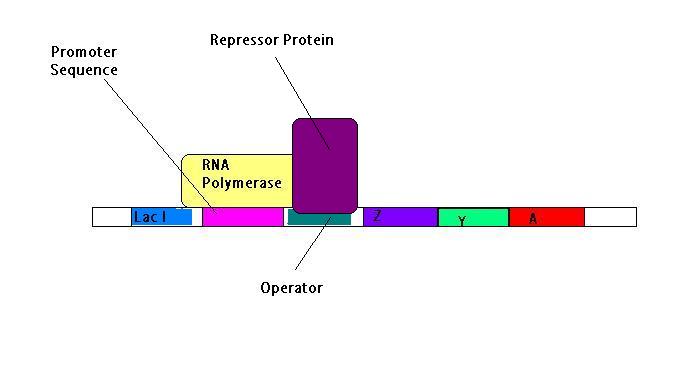
The repressor protein is a homotetramer and a product of the lacI gene, and will bind tightly to the operator, under the correct conditions i.e. when glucose is present and lactose absent. When the repressor is bound to the operon, the RNA polymerase is unable to unwind the DNA in order to expose the bases and hence is unable to transcribe the structural genes as there is no template for the RNA synthesis to occur. The group of structural genes act as a single transcription unit, coding for a single mRNA molecule termed a polycistronic transcript i.e. coding for multiple proteins and transcription is dependent upon the correct environmental conditions as described below:
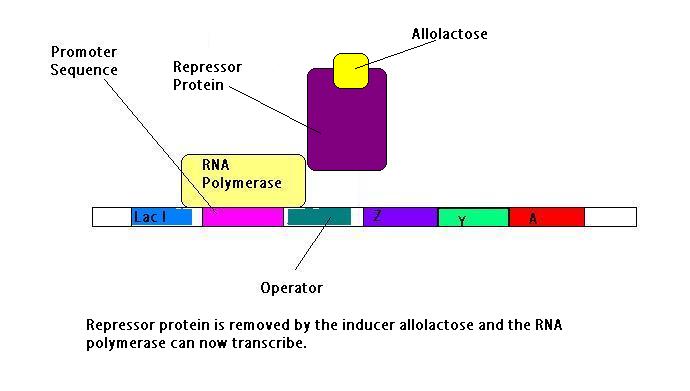
In the presence of lactose, the lac operon is activated by allolactose. Lactose is converted to allolactose by the enzyme beta-galactosidase. This conversion s essential as allolactose binds to the lac repressor and a conformational change occurs - it shifts the formation of the DNA binding subunits by 3.4A, this reduces the affinity for the operator by 1000 times.
In the absence of glucose, cAMP accumulates (this is because glucose metabolites inhibit adenylate cyclase and therefore prevent cAMP formation). cAMP is important because it controls the activity of an activator protein called CAP. When CAP is bound to cAMP, it binds to a binding site on the lac operon thereby, activating transcription[2].
What is the Lac operon?
The lac operon is a good example of how genes are regulated, in this case through the acts of an activator and/or repressor.The lac operon was studied in E. coli[3]. It contains 3 genes that are needed to produce proteins that are required to break down lactose when it is present in the cell. These 3 genes are Lac Z, Lac Y and Lac A. Each code for B- galactosidase (an enzyme which hydrolyses lactose into glucose and galactose), Permease (an enzyme which promotes lactose transport across the cell membrane of bacterium) and Transacetylase (an enzyme which transfers acetyl groups from acetyl-coenzyme A to B--galactoside), respectively[4].
Further up the genetic code from these three genes, upstream, lies the promoter sequence. RNA polymerase needs a region in which it can join the genetic code, the Promoter sequence, before it can start transcribing. RNA polymerase is required in transcription of the Lac operon[5]. 
When does the Lac operon function?
The Lac operon is regulated depending on the absence or presence of certain substances. When lactose is present in the cell and glucose is absent, then the Lac operon is active and the 3 genes are transcribed to break down this lactose in the cell.
The lac operon only needs to activate the genes necessary for lactose integration when there is an absence of glucose and a plentiful supply of lactose.
Negative gene regulation
The conditions inside the cell are changing all the time. So what happens when glucose is present and lactose levels are low? The Lac operon is no longer required to make the proteins to break down lactose and so its function is switched off. This is done by the use of a repressor protein[6].
Upstream of the promoter sequence there is another gene. This is the Lac I gene. The Lac I gene is transcribed to make the repressor protein which binds to 3 different operator sequences present at different parts of the Lac Operon. The repressor protein is a tetramer and the binding of this repressor to the operators cause the DNA to be looped around.
Once the repressor protein is bound, it stops the RNA polymerase enzyme from transcribing the genes thus effectively it acts as block[7].
When the glucose levels are depleted and the lactose levels are high, the repressor has to be removed in order to transcribe the required genes. This is done by an inducer molecule. This molecule comes from lactose and is Allolactose. Allolactose is synthesised from lactose by the enzyme B-galactosidase, which is also a translational product of the lac gene. The allolactose binds to the repressor protein and causes a conformational change in the repressor which causes it to dissociate from the DNA.Upon binding the allolactose reduces the affinity of the repressor protein for the lac operator by a factor of 1000. RNA polymerase can work as it is not blocked and the Lac Z, Lac Y and Lac A genes are transcribed[8].
Positive Gene regulation
Sometimes promoters are not strong enough to initiate transcription on their own and so require another molecule or complex to help. In the Lac operon, this is done by the CRP – cAMP complex. This is because glucose acts as an inhibitor of the enzyme adenylyl Cyclase which is responsible for converting ATP into cAMP (cyclic adenosine monophosphate)When glucose levels in the cell are low, the levels of cAMP increase as glucose no longer blocks cAMP synthase[9]. This then combines with a CRP protein (sometimes also referred to as the CAP protein) and forms a complex called the CRP-cAMP complex; CRP can activate transcription at more than 100 catabolite-sensitive operons. Thus, this complex then joins to a sequence of nucleotides downstream from the LacI promoter known as the CRP binding site. The binding increases the affinity of RNA polymerase for the lac promoter sequence and hence it binds and as a result, transcription can take place[10].When the levels of glucose increase again, the amount of cAMP synthesised is reduced and so the complex levels decrease. This, therefore, inhibits the Lac operon from working because the affinity of the RNA pol II for the lac gene promoter without the activator bound is quite low.
References
- ↑ Cellular and Molecular Life Sciences, Matthews KS, Swint-Kruse L, Wilson CJ, Zhan H, The Lactose Repressor System: Paradigms for Regulation, Allosteric Behaviour and Protein Folding.January 2007; 64(1):3-16
- ↑ Biochemistry pgs.963-964, 7th edition, J M.Berg, J L. Tymoczko, L. Stryer
- ↑ Hartl, D. L. and Jones, E. W., 2009. Genetics: Analysis of genes and genomes. 7th edition. Sudbury: Jones and Bartlett Publishers pg 383
- ↑ http://users.rcn.com (2011) "The operon" – 30th March 2010 – Available from: http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/L/LacOperon.html [Accessed 3rd January 2011]
- ↑ Hartl, D. L. and Jones, E. W., 2009. Genetics: Analysis of genes and genomes. 7th edition. Sudbury: Jones and Bartlett Publishers pg 386
- ↑ Hartl, D. L. and Jones, E. W., 2009. Genetics: Analysis of genes and genomes. 7th edition. Sudbury: Jones and Bartlett Publishers pg 384
- ↑ Sadava (2011) “ The Lac Operon” – 2008 – Available from: http://www.sumanasinc.com/webcontent/animations/content/lacoperon.html [Accessed 3rd January 2011]
- ↑ BioCoach Activity (2011) "The lac inducer: Allolactose" – Available from: http://www.phschool.com/science/biology_place/biocoach/lacoperon/inducer.html [Accessed 3rd January 2011]
- ↑ Mulligan, M. E. (2002) “The lac operon: positive regulation” – Available from: http://www.mun.ca/biochem/courses/3107/Topics/Lac_positive_control.html [Accessed 3rd January 2011]
- ↑ Hartl, D. L. and Jones, E. W., 2009. Genetics: Analysis of genes and genomes. 7th edition. Sudbury: Jones and Bartlett Publishers pg 389