Riboswitches
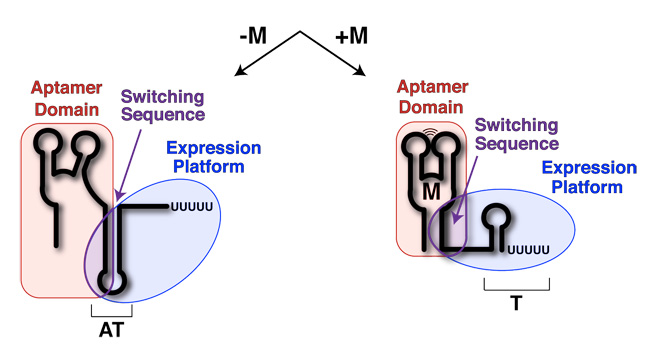
Riboswitches are short sequences of RNA, found near the 5' end of mRNAs, before the start codon. They are able to bind metabolites and other small molecules, acting as ligands, to bring about a change in there confirmation. This conformation, due to the complex formed with the complimentary ligand, goes on to regulate gene expression. The use of the complex is to act as a control against gene expression. When the mRNA is being synthesised, the riboswitch sequence folds and will only allow progress of the RNA polymerase if the regulatory ligand hasn't formed a complex with the riboswitch sequence.[1]
The use riboswitches is prolific in bacteria. A main use of riboswitches by bacteria is regulation of ribonucleic sequences, encoded by the cis acting elements in the full transcript. Further to this, it is now known that riboswitches are not always found at the 5' end, as seen in some eukaryotic mRNA. Thiamine Pyrophosphate riboswitch is found at the 3' end and is known to regulate splicing..[2]
References
- ↑ Alberts, B., Johnson, A., Lewis, J., Raff, M., Roberts, K. and Walter, P. (2008)Molecular Biology of the Cell, 5th edition, New York: Garland Science.
- ↑ http://www.nature.com/scitable/topicpage/riboswitches-a-common-rna-regulatory-element-14262702